 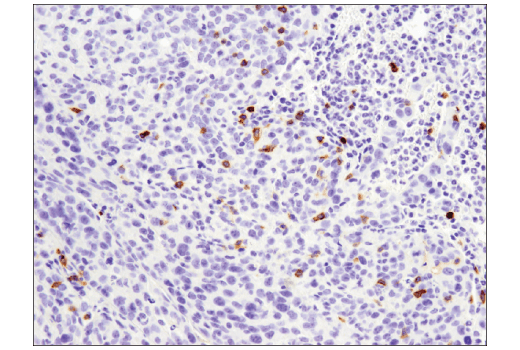 
This subgroup of proteins are of particular interest for basic and applied research due to their unique signaling functions, enabling, limiting and orchestrating cellular communication and interactions. The surfaceome represents the subgroup of proteins at the plasma membrane with exposed domains towards the extracellular space including for example G-protein coupled receptors, receptor tyrosine kinases and integrins. However, such information is a prerequisite to understand concerted biological functions of cell types in complex signaling environments. Although knowledge about molecular features of these cell types is gathered at ever increasing speed, detailed information about the expressed cell surface protein repertoire of individual cell types is sparse due to technological limitations. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist.Īccording to traditional phenotypic classification systems, the human body contains approximately 210 functionally distinct cell types. Program in Molecular Life Sciences in Zurich (AH) Neuroscience Center Zurich Fellowship, (TB) Interdisciplinary PhD Project of SystemsX.ch, the Swiss Initiative of Systems Biology, Project 2 2009, (HM) MD-PhD fellowship of the Swiss National Science Foundation, (FC) Advanced European Research Council grant #233226, (RA) Swiss National Science Foundation (grant # 3100A0-107679), (RA) National Institutes of Health (NIH Grant # U01CA152813), (RA). 
The work is made available under the Creative Commons CC0 public domain dedicationĭata Availability: The MS-based proteomics data have been deposited to the ProteomeXchange Consortium ( ) via the PRIDE partner repository with the dataset identifier PXD000589.įunding: This study was supported by the National Center of Competence in Research Neural Plasticity and Repair of the Swiss National Science Foundation, Center for Proteomics, (BW) InfectX and BIP project within the Swiss Initiative in Systems Biology, (BW) Mueller Fellowship from the Ph.D. This is an open access article, free of all copyright, and may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose. Received: NovemAccepted: JanuPublished: April 20, 2015 PLoS ONE 10(4):Īcademic Editor: Hong Wanjin, Institute of Molecular and Cell Biology, UNITED STATES (2015) A Mass Spectrometric-Derived Cell Surface Protein Atlas. 
The CSPA will be useful for the evaluation of drug targets, for the improved classification of cell types and for a better understanding of the surfaceome and its concerted biological functions in complex signaling microenvironments.Ĭitation: Bausch-Fluck D, Hofmann A, Bock T, Frei AP, Cerciello F, Jacobs A, et al. Integrated analysis of the CSPA reveals that the concerted biological function of individual cell types is mainly guided by quantitative rather than qualitative surfaceome differences. The cellular surfaceome snapshots of different cell types, including cancer cells, resulted in a combined dataset of 1492 human and 1296 mouse cell surface glycoproteins, providing experimental evidence for their cell surface expression on different cell types, including 136 G-protein coupled receptors and 75 membrane receptor tyrosine-protein kinases. The CSPA is presented in form of an easy-to-navigate interactive database, a downloadable data matrix and with tools for targeted surfaceome rediscovery ( ). Here, we applied the Cell Surface Capture (CSC) technology to 41 human and 31 mouse cell types to generate a mass-spectrometry derived Cell Surface Protein Atlas (CSPA) providing cellular surfaceome snapshots at high resolution. However, information about the cell surface protein repertoire (the surfaceome) of individual cells is only sparsely available. Cell surface proteins are major targets of biomedical research due to their utility as cellular markers and their extracellular accessibility for pharmacological intervention. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
